Study of host-viral interactions of Crimean-Congo haemorrhagic fever identified potential new drugs
The COVID-19 pandemic is now moving into a more endemic phase in Sweden, as well as many other parts of the world. Regardless, SARS-CoV-2 still poses a serious global health threat, particularly when considering the potential effects of novel variants. Continuous pandemic preparedness efforts are therefore warranted. Global preparedness is also beneficial to safeguard against the threat of pandemics caused by other emerging pathogens.
At present, Crimean-Congo hemorrhagic fever virus (CCHFV) is uncommon in most parts of the world. However, it is endemic to almost 30 countries across sub-Saharan Africa, South-Eastern Europe, the Middle East, and Central Asia (Bente et al., 2013; Zivcec et al., 2016), and is spreading into new populations. CCHFV has been capable of infecting humans for thousands of years; archeological blood samples indicate that it was endemic to Celtic Iron Age populations near the Danube river. CCHFV is an RNA virus that belongs to the Nairoviridae family, and causes Crimean-Congo hemorrhagic fever (CCHF) disease. CCHF is transmitted through bites from infected ticks and can pass from animals to humans as well as between humans. Symptoms of the disease include fever, joint pain, vomiting, and uncontrolled bleeding, which can ultimately lead to organ failure. To date, there are no effective treatments for or vaccine against CCHF, and mortality rates for the disease vary between 3 and 40%.
Knowledge about the pathogenesis of CCHFV, and how it interacts with host cells is key to enable future drug and vaccine development. However, very little is currently known, primarily because outbreaks of CCHF are sporadic and usually occur in regions lacking the infrastructure to conduct the necessary studies.
Recently, a large study looking at the pathogenesis and host-viral interactions of the CCHFV has been published (first author: Ujjwal Neogi; corresponding author: Ali Mirazimi). The work involved researchers from multiple Swedish institutions, including Karolinska Institutet, the Public Health Agency, Stockholm University, Bioinformatics Infrastructure Sweden (NBIS), Science for Life Laboratory (SciLifeLab), and the National Veterinary Institute of Sweden. They were joined by research groups from India, Turkey, Germany and the UK, making this a truly international effort.
Neogi and colleagues used a multi-omics systems biology approach to study the complex host-viral interaction of the CCHFV. The same approach was recently used by the team to study the response of other viruses e.g. HIV-1, SARS, MERS and SARS-CoV-2, (Appelberg et al., 2020; Krishnan et al., 2021). We previously wrote a Data Highlight about some of their earlier work on the Swedish COVID-19 & Pandemic Preparedness Data Portal. In that study, the researchers aimed to mimic the alternations that occur during the acute CCHFV viral phases.
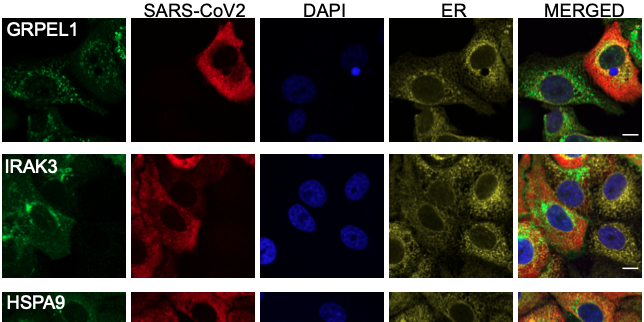
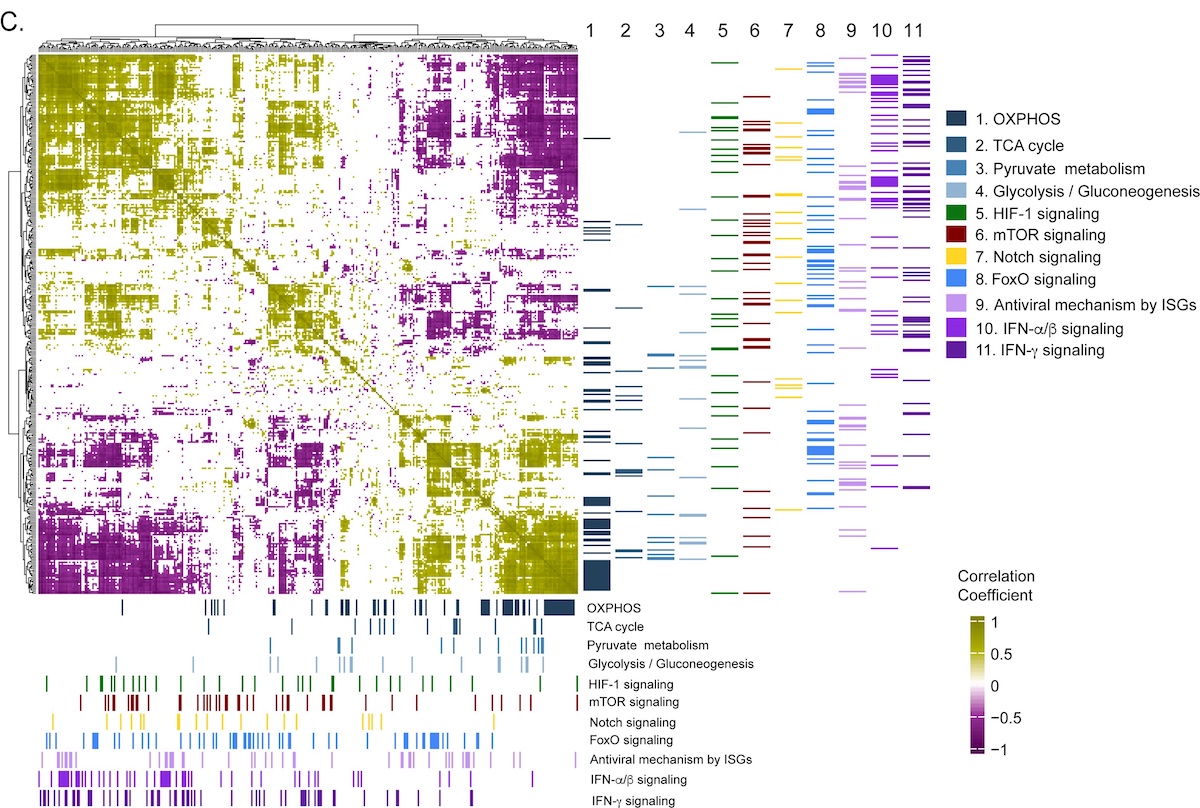
In the present study, the researchers completed a longitudinal study of a CCHFV patient cohort originating from Turkey (initial N=18, mortality rate 5.6%, 12 were included in the study). They analyzed peripheral blood mononuclear cells (PBMCs) during both the acute and convalescent (after one year of recovery) phases of the disease. They used a differential gene expression (DGE) profile to compare gene expression in the two phases. This showed that 2891 genes were up-regulated, and 2738 genes were down-regulated. The researchers then used RNAseq analysis of PBMCs in order to compare the metabolic pathways used in the acute and convalescent phases. In brief, the results showed a number of changes (N=205), with ~30% up-regulated and ~10% down-regulated in the interferon signaling-related pathways, which are known to be involved in viral response during CCHFV infection. In addition, pathways involved in central carbon and energy metabolism (CCEM) were found to be important during the CCHFV infection. Notably, an up-regulation of oxidative phosphorylation (OXPHOS) was detected in both transcriptomics and Huh7 proteomics data from patients. In addition, a time-series analysis of a Huh-1 cell culture confirmed that OXPHOS was increasingly up-regulated during the productive viral replication phase. The researchers then directly tested how viral replication was affected by blocking two key CCEM pathways fueling OXPHOS in the Huh7 cells; glycolysis and glutaminolysis. Specifically, they used 2-deoxy-D-glucose (2-DG) and 6-diazo-5-oxo-L-norleucine (DON), both of which are known drug candidates. This resulted in suppressed CCHFV-replication and successfully controlled viral replication in the lab setting.
“Our main goal to take this approach targeting host to limit the viral replication subsequently disease severity to the clinic, and we are initiating the pre-clinical studies in animal experiments.” said Ujjwal Neogi, professor at the Department of Laboratory Medicine, Karolinska Institutet.
In summary, the findings in this article could represent the first step towards the new antiviral drug treatment against CCHF disease. It is clear that further studies into the CCHF pathogenesis are required to investigate the host-virus interactions of CCHF. However, this study showed that targeting OXPHOS, and increasing the host antiviral response, may be one way to develop an attractive host-directed therapy during the acute CCHFV infection. CCHFV has been infecting human populations for thousands of years, and remains rare globally. However, CCHF is already an endemic in some parts of the world and continues to spread. This, taken together with the fact that no effective vaccines or treatments available for the disease, means that it poses a serious risk to human health that cannot be ignored. The development of treatments and vaccines against infectious diseases for which none are currently available is an important part of pandemic preparedness efforts.
Data
- Raw RNAseq data is available in Sequence Read Archive (SRA) with ID PRJNA680886.
- Mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD022672.
- Analysis code is available on GitHub.
Article
Neogi, U., Elaldi, N., Appelberg, S., Ambikan, A., Kennedy, E., Dowall, S., Bagci, B. K., Gupta, S., Rodriguez, J. E., Svensson-Akusjärvi, S., Monteil, V., Vegvari, A., Benfeitas, R., Banerjea, A., Weber, F., Hewson, R., Mirazimi, A. (2022) Multi-omics insights into host-viral response and pathogenesis in Crimean-Congo hemorrhagic fever viruses for novel therapeutic target. eLife, 11, e76071. 10.7554/eLife.76071.
Funding
This study was supported by grants from the Swedish Research Council, the Public Health England Grant In Aid, Karolinska Institutet and the European Union Horizon 2020 CCHF Vaccine Grant.